The Frustrations
Medical image annotation is full of operational frictions, to name a few:
- MANAGING data/datasets is hard: as I mentioned in the challenge of medical image analysis, a huge data space, many sources, and varying quality
- MANAGING annotations is also hard: you have a team of annotators with different expertise and styles, you need to align their ability AND style while doing multiple rounds/versions
- MANAGING tasks is also hard: you can’t label everything AT ONCE, it’s too laborious and impossible to review, but you eventually need to piece them together.
These are real problems, but I don’t see them as the fundamental challenge of our field. They are frustrations—matters of process and diligence that can be solved with better tools and project management.
The Challenge
Unlike many computer vision tasks where a label can be applied with common sense, medical imaging operates on a different plane. It presents two distinct types of work, each demanding a different skill set:
- Laborious Delineation: This is the meticulous, often tedious work of finding minuscule lesions or outlining structures. Think of scrolling slice-by-slice through a CT scan to find every “blob-like” lung nodule or painstakingly segmenting the intricate network of liver vessels. This work demands immense patience, focus, and dexterity.
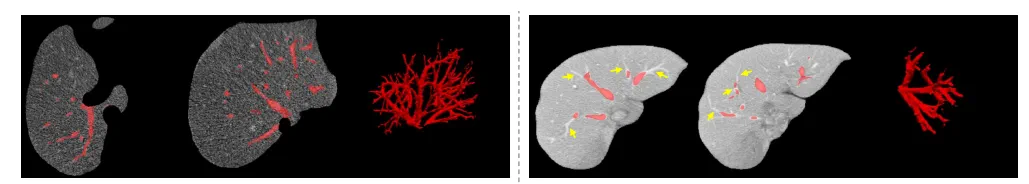
Illustration of liver vessels. Left: what you want to label; right: what you start with from a model prediction. Figure from a paper for a different context

- Expert-Driven Diagnosis: This is the detective work of clinical interpretation. It involves identifying a subtle, unusual abnormality, understanding its context, and performing a differential diagnosis. This requires deep clinical knowledge—knowing the potential suspects, their patterns, and their anti-patterns—that takes years to build.
The central problem arises when we misalign these tasks and the people performing them. It’s wasteful to have a trained radiologist spend hours on the laborious work of segmenting vessels. It’s futile—and dangerous—to have a non-expert attempt the expertise-driven work of diagnosing a rare condition.
You might argue: why not just hire an all-expert annotation team?
My own experience has shown this is not only uneconomical but often yields suboptimal results for delineation tasks. Experts, rightly focused on diagnostic subtleties, may have different “styles” of segmentation that are difficult to standardize.
Unsurprisingly, sometimes it is the annotators WITHOUT clinical expertise who produce better-quality annotations due to the sheer amount of attention and energy they invest.
The Solution
The answer to this expertise allocation problem is to divide and conquer.
We must separate the concerns of “what does it look like?” (delineation) from “what is it?” (diagnosis).
A straightforward approach is to have experts perform a quick triage. They can rapidly review cases, flagging abnormalities and narrowing the data space. This allows a team of trained annotators to focus their efforts on the laborious delineation of these pre-identified regions.
An even better, more scalable solution is to build a bootstrapping system of models. Here, different models have different responsibilities. An initial detection or segmentation model can perform the first pass. It doesn’t have to be perfect. Its role is to handle the 80% of laborious work, flagging regions of interest for human review. This frees up experts to focus on the most critical and ambiguous cases, correcting the model’s mistakes and providing the high-level diagnostic labels. This human-in-the-loop system allows for greater flexibility and maximizes the value of every minute of an expert’s time.
Final Thought
This brings me to a final, crucial point about the very nature of annotation.
In traditional computer vision, we “transcode” rich visual information into simple numeric labels or masks. This process strips the data of its original context, forcing models to learn from mapping a high-dimensional image to a sparse label index. Humans, in contrast, learn from far fewer examples because we learn from much richer signals, not just simple labels.
This suggests a paradigm shift. What if we could “teach” our models with more semantic annotations? Instead of just outlining a tumor, what if the annotation included notes like “spiculated margins” or “adjacent to a vessel”? By enriching our labels with clinical reasoning, we might build models that learn more efficiently, perform more robustly, and require significantly less data.
The future of medical annotation may not be just about creating more labels, but about creating smarter ones.
