Trust me, I DO have conviction in the scaling law—the paradigm that data + compute is all you need.
I find the “less structure, more intelligent” 2 idea very appealing.
BUT, we are just not there yet, at least in the medical imaging domain.
So, here we are, leveraging prior knowledge (aka domain knowledge) to boost training efficiency and data utilization.
Before I dive into medical imaging specifics, I would like to point out:
Many natural imaging CV tasks leverage prior knowledge as well
- I would argue that pose landmark detection relies on a HUMAN-DEFINED interpretation of skeletal structure , instead of a naturally learned representation.
- BEV feature aggregation is predefined by human-calculated epipolar consistency.
- Speaking of autonomous vehicles, this target trajectory prediction paper on modeling the coordinate system in transformer positional embedding is pretty interesting 1 .
Radiologists utilize prior knowledge to build better visualizations
-
Curved multiplanar reformation (CPR) and stretched multiplanar reformation (sMPR) are commonly used to diagnose coronary disease. These techniques essentially stretch the 3D vessel along a plane into a straight line for easier interpretation of its structure and interior by eliminating other information from vast 3D voxels.
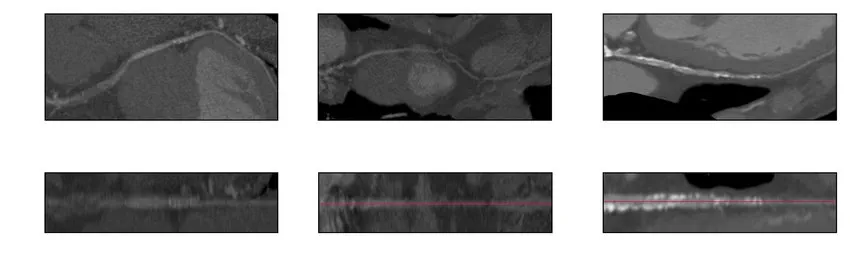
-
Similarly, rib-unfolding visualizations make it easier and faster to pinpoint rib fractures, even for non-experts.
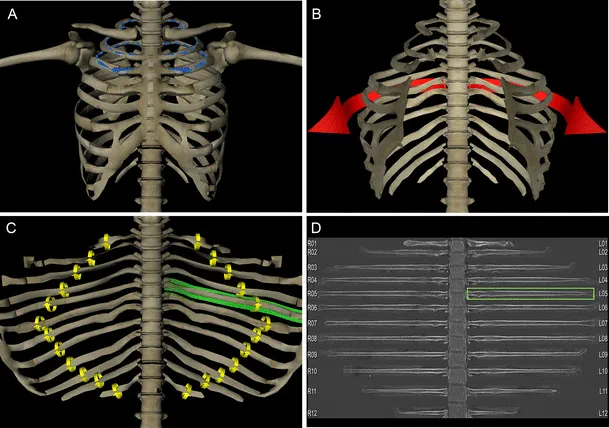
Examples in medical imaging analysis
There are MANY more—I just name a few I’ve encountered over the years
-
Mass detection in mammography: mass features should be correlated across multiple views
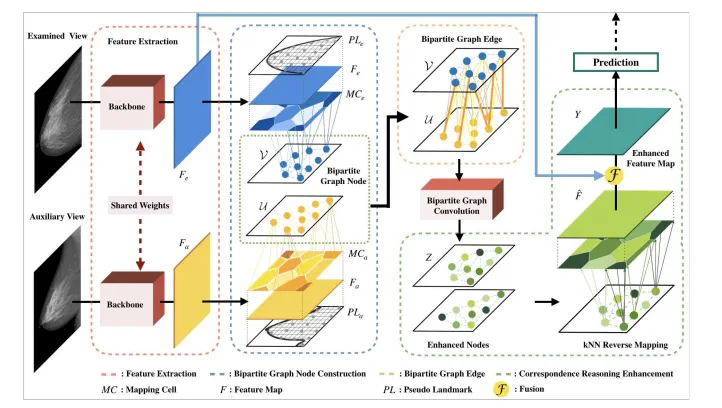
-
Vertebra localization and identification in CT: vertebrae should form a coherent centerline along the spine
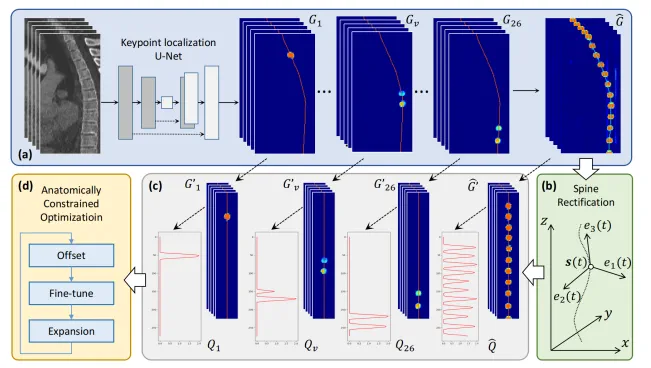
-
Suspicious node malignancy prediction: should account for surrounding lymph node conditions
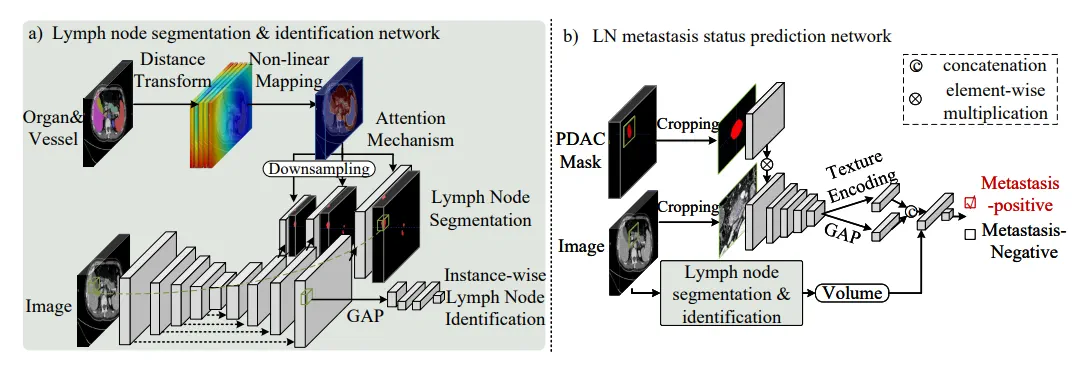
-
Vessel centerline extraction: easier/better to model with graphs (nodes) than with segmentation (voxels)
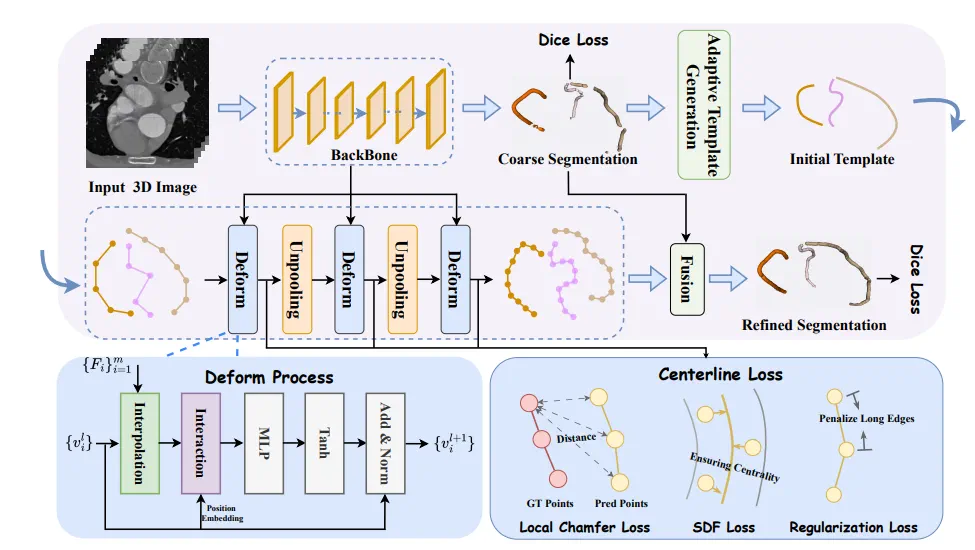
DeformCL: Learning Deformable Centerline Representation for Vessel Extraction in 3D Medical Image
Not to mention, algorithms also build on top of specialized reconstructions for human interpretation. Unsurprisingly, what’s easier for radiologists to tell is also easier for models to learn.
What is leveraging prior knowledge anyway?
Essentially, leveraging prior knowledge is the act/technique of carrying biologically/physically/pathologically grounded knowledge into the model, in the form of “tensor affinity”:
- Choose a level of abstraction: pixel space, spatial feature space, semantic feature space, instance feature space, etc.
- Heuristically set “communication rules” (which tensors should communicate with which)
- Choose a method of communication (feature aggregation) - transformer is the default
- Note the result might be warped into a whole new space , e.g., dense pixels → sparse nodes, and vice versa
Prior knowledge in medical imaging
The most important prior of the human body we should leverage is the anatomical prior:
- Landmarks: Key anatomical reference points
- Relative stability: Position and shape relationships between body parts remain relatively stable
- Symmetry: Bilateral symmetry in many structures
- Topological structure: I find this particularly interesting but under-explored. Most approaches assume oversimplified topology and utilize mathematical tools like Betti numbers to model it
The second most important prior is the human biological system:
- Diseases affecting one biological system often impact multiple parts of that system concurrently
